Spine Segmentation | Master Thesis
Anatomical labeling of vertebrae and intervertebral discs (IVDs) in small field-of-view MRI scans.
PyPI · Master's Thesis (PDF) · Slides · GitHub
This library provides pretrained deep learning models for segmenting and labeling individual vertebrae (C2 to S1) and IVDs in 3D MRI volumes. It works on small field-of-view scans where only part of the spine is visible -- the common case in clinical practice, since full-spine MRIs are expensive and time-consuming.
The approach was developed as part of a master's thesis (submitted November 2023) and achieves 85.5% subset accuracy when only ~20% of the spine is visible, improving to 92.6% at ~40% and 94.4% at ~60% visibility.
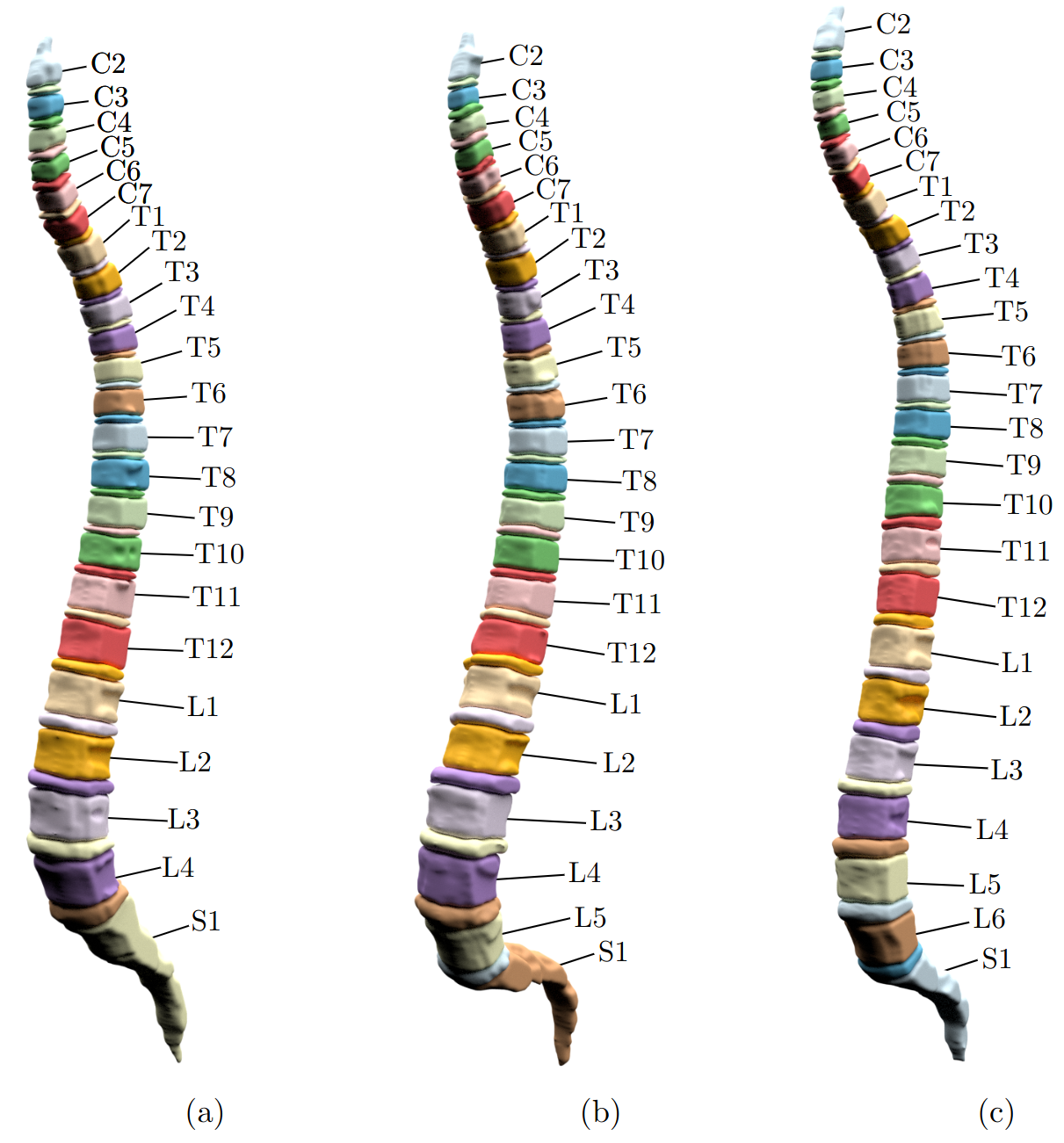
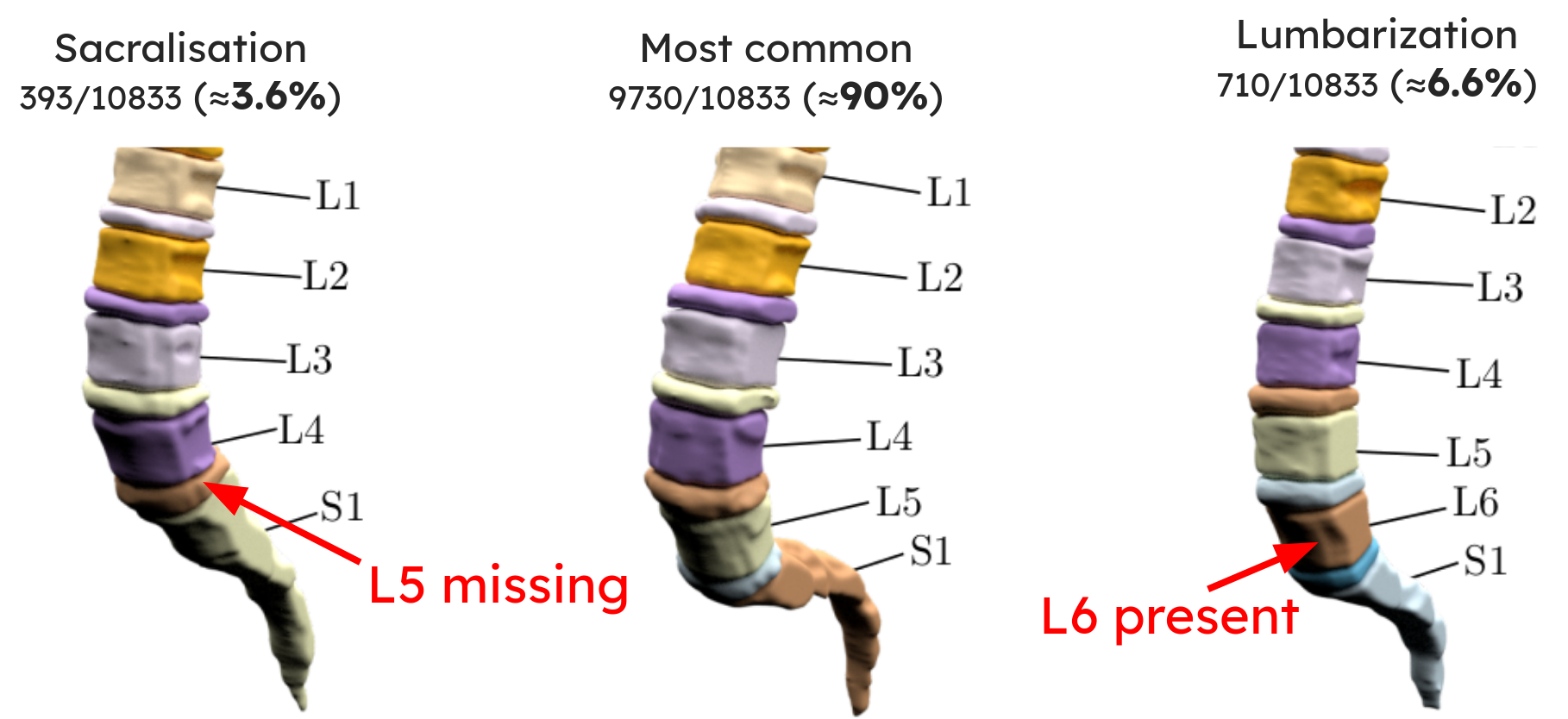
Above: three spines automatically segmented and labeled. (a) shows sacralization (L5 fused with sacrum, only 4 lumbar vertebrae), (b) a normal spine, (c) lumbarization (additional vertebra L6).
Installation
pip install spine-segmentation
Usage
The input should be a 3D numpy array representing an MRI volume (shape: slices x width x height). The library handles resizing internally.
import numpy as np
from spine_segmentation import SegmentationInference
mri_volume = ... # shape: (num_slices, width, height)
inference = SegmentationInference(
segmentation_device="CPU", # or "GPU"
instance_segmentation_device="CPU", # or "GPU"
output_same_shape_as_input=True, # pad/crop output to match input shape
)
result = inference.segment(mri_volume)
result.semantic_segmentation # 3D array: 0=background, 1=vertebra, 2=IVD
result.instance_segmentation # 3D array: each vertebra/IVD has a unique integer ID
result.id_to_label # dict mapping ID -> anatomical label (e.g. {1: "C2", 2: "C2-C3", ...})
For 2D slices, repeat the slice three times to create a minimal 3D volume:
mri_volume = np.tile(single_slice, (3, 1, 1))
You can also run only the 2-class segmentation (vertebra vs. IVD, no instance labeling) which is faster:
result = inference.segment(mri_volume, only_2class_segmentation=True)
Models are downloaded automatically on first use.
How it works
The pipeline has three steps:
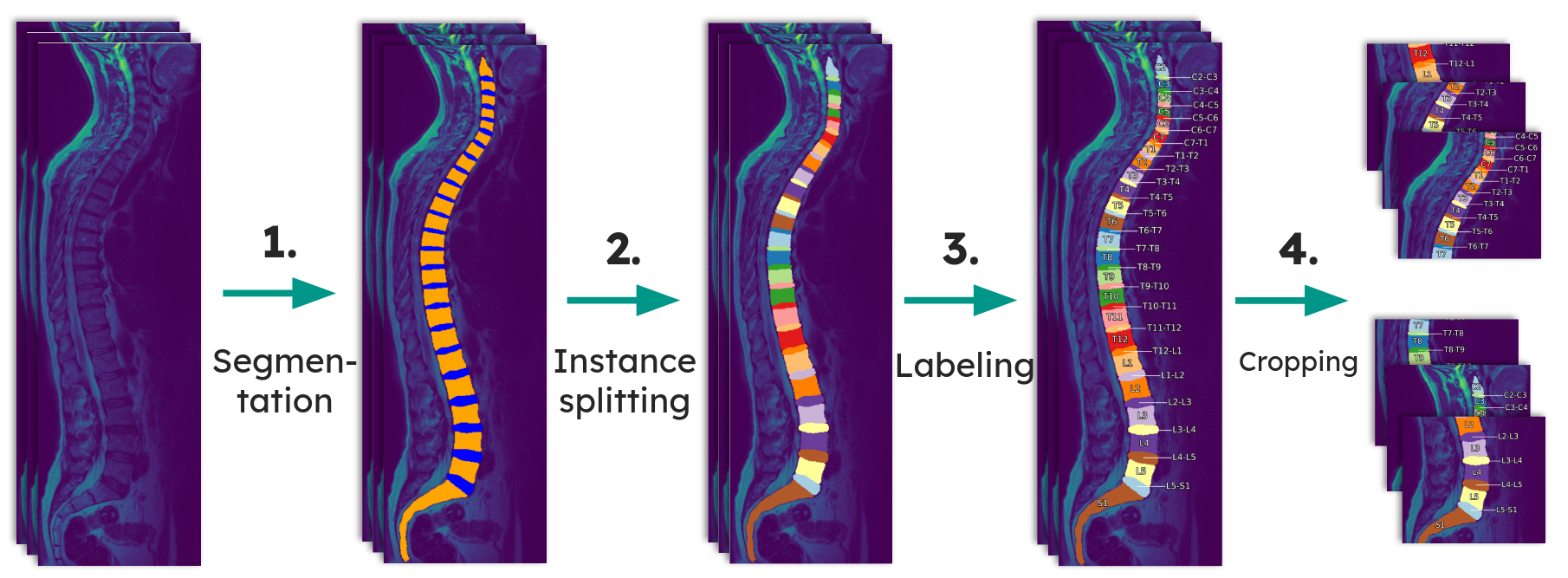
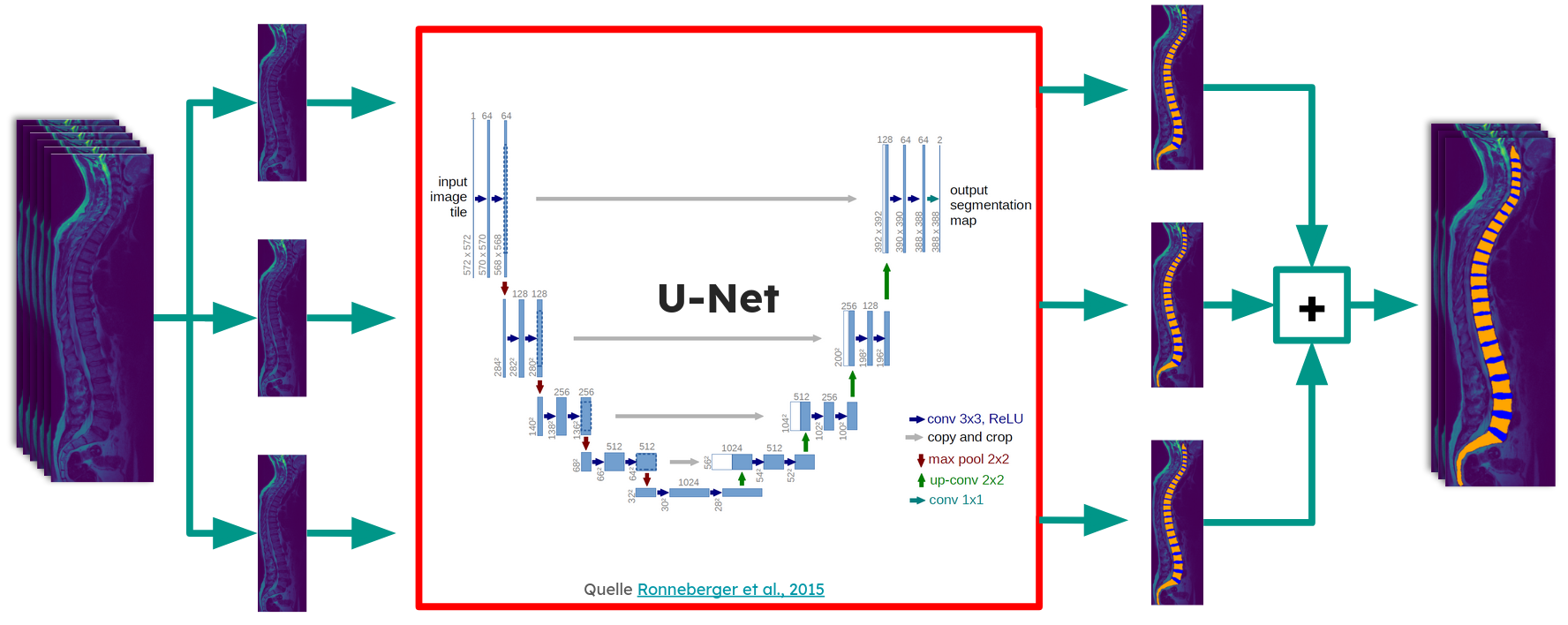
Segmentation -- A slice-wise 2D U-Net (ResNet152 encoder, Jaccard loss) segments each sagittal slice into vertebra, IVD, and background. The best model achieved a Dice score of 0.919 on the validation set, outperforming a prior 3D patch-based approach on the same dataset. Each slice is fed through the network together with its two adjacent slices (as RGB channels), and the resulting 2D segmentations are reassembled into a 3D volume.
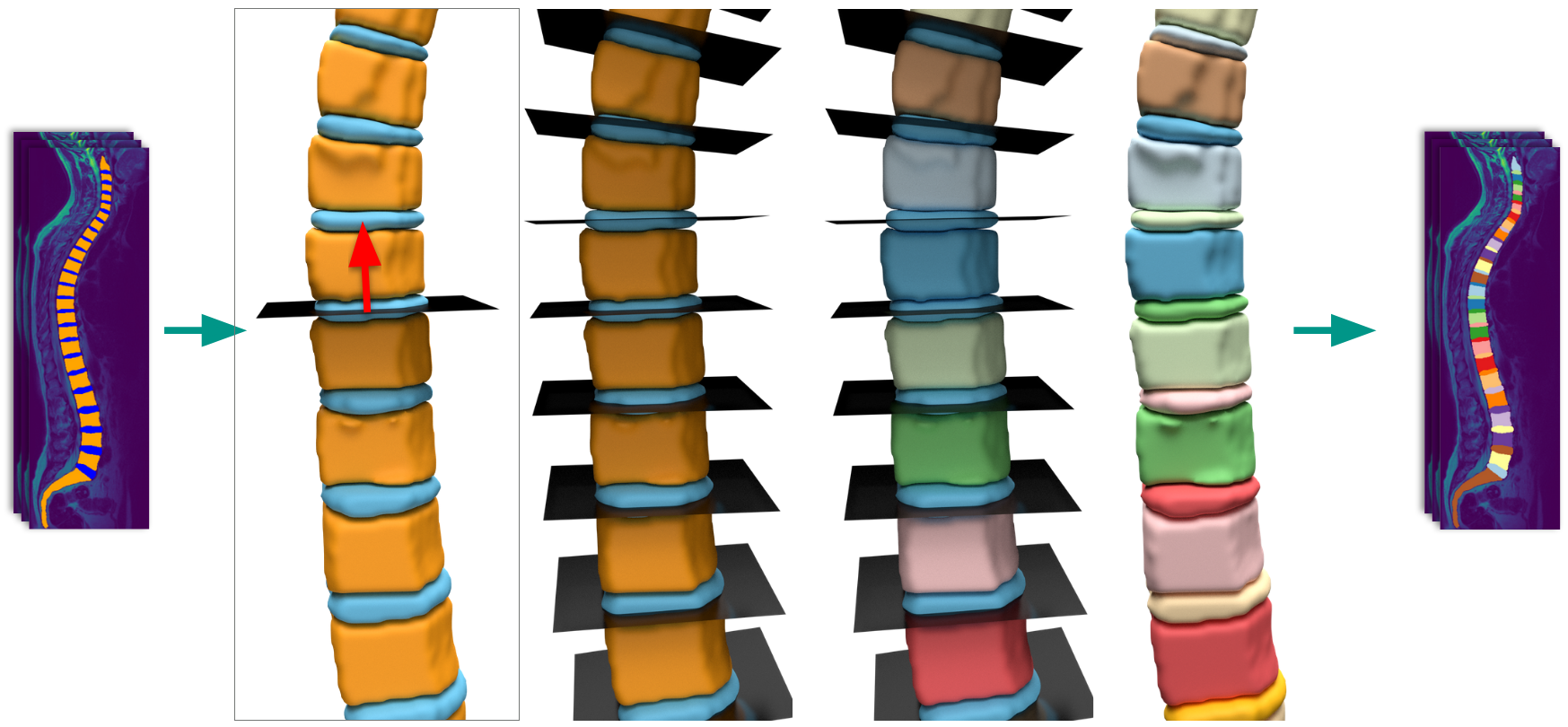
Instance splitting -- Connected vertebra/IVD regions are separated into individual instances. The best method splits along the IVD boundaries using plane fitting, achieving 99.2% valid separations on 10,833 scans.
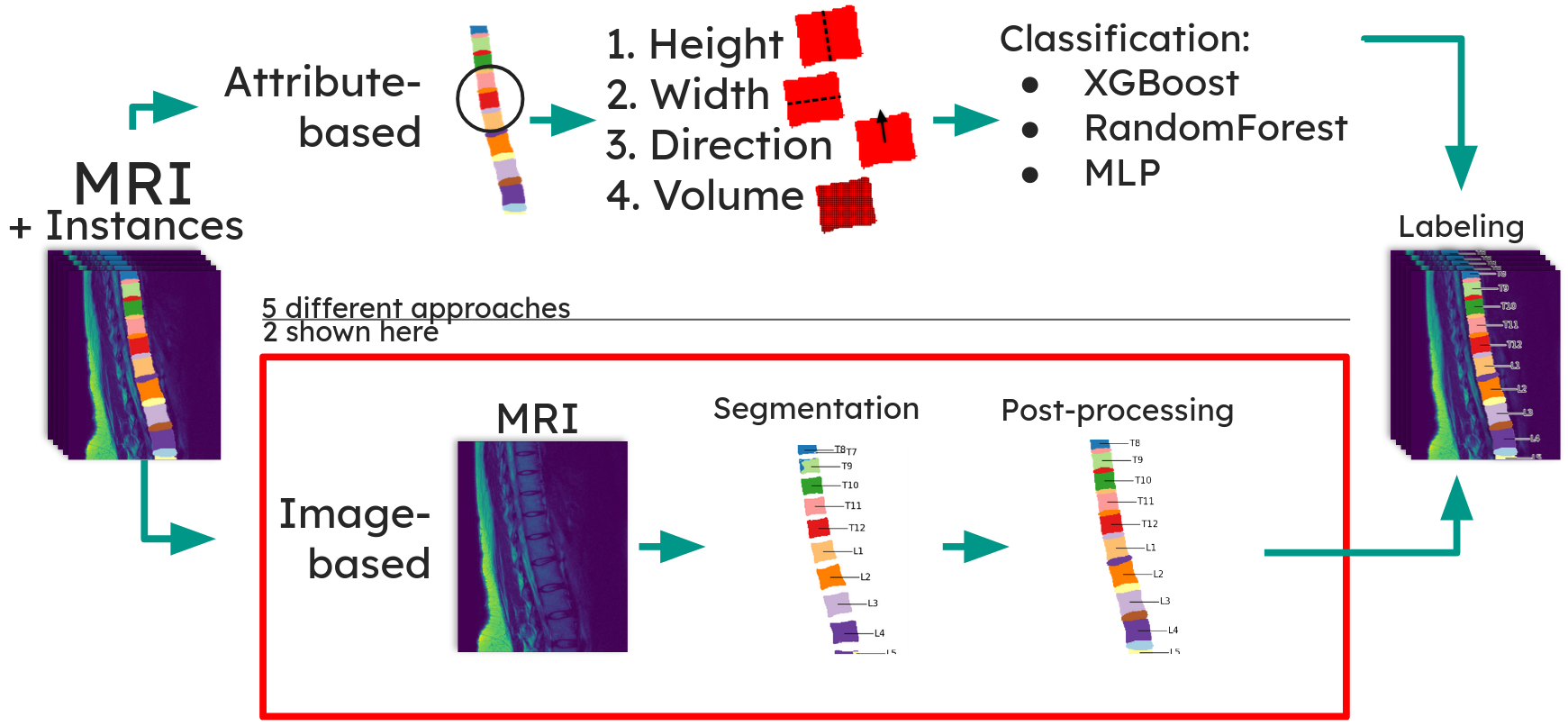
Anatomical labeling -- A multiclass segmentation model assigns the correct anatomical label (C2, C2-C3, C3, ..., S1) to each instance. Five approaches were evaluated; a direct multiclass U-Net segmentation trained on automatically generated ground truth performed best.
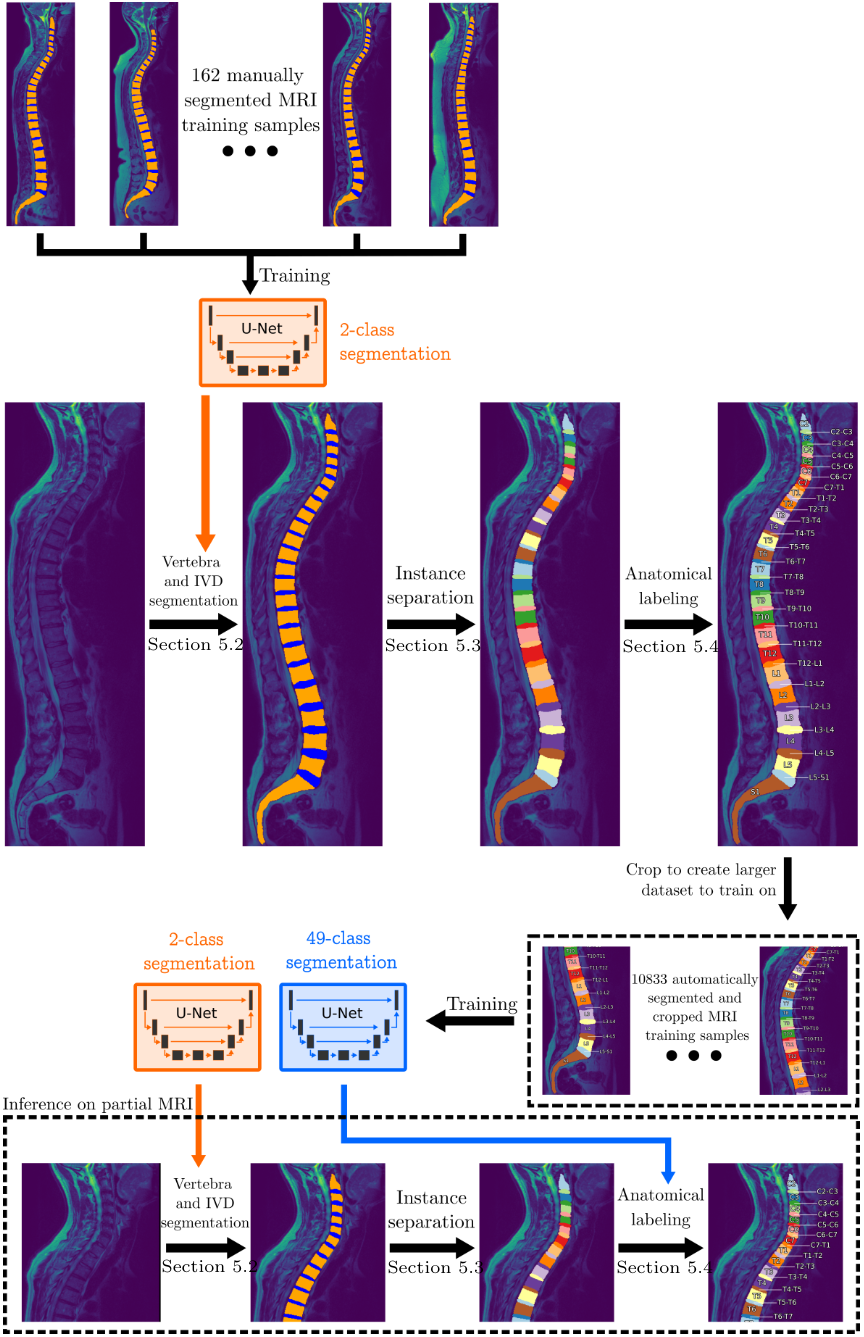
The key insight enabling this pipeline is that a large dataset of 10,833 complete MRI scans can be used to generate training data for small-FOV labeling. A model trained on 162 expert-annotated scans first segments the full dataset. Since the topmost vertebra in a complete scan is always C2, all labels can be inferred automatically. Cropped subsets of these labeled scans then serve as training data for the small-FOV labeling model.
Key results
Subset accuracy (every vertebra and IVD in the scan must be labeled correctly):
| Visible vertebrae | ~% of spine | Subset accuracy | Dice score |
|---|---|---|---|
| V=5 | ~20% | 85.5% | 0.799 |
| V=8 | ~30% | 90.9% | 0.839 |
| V=10 | ~40% | 92.6% | 0.847 |
| V=15 | ~60% | 94.4% | 0.875 |
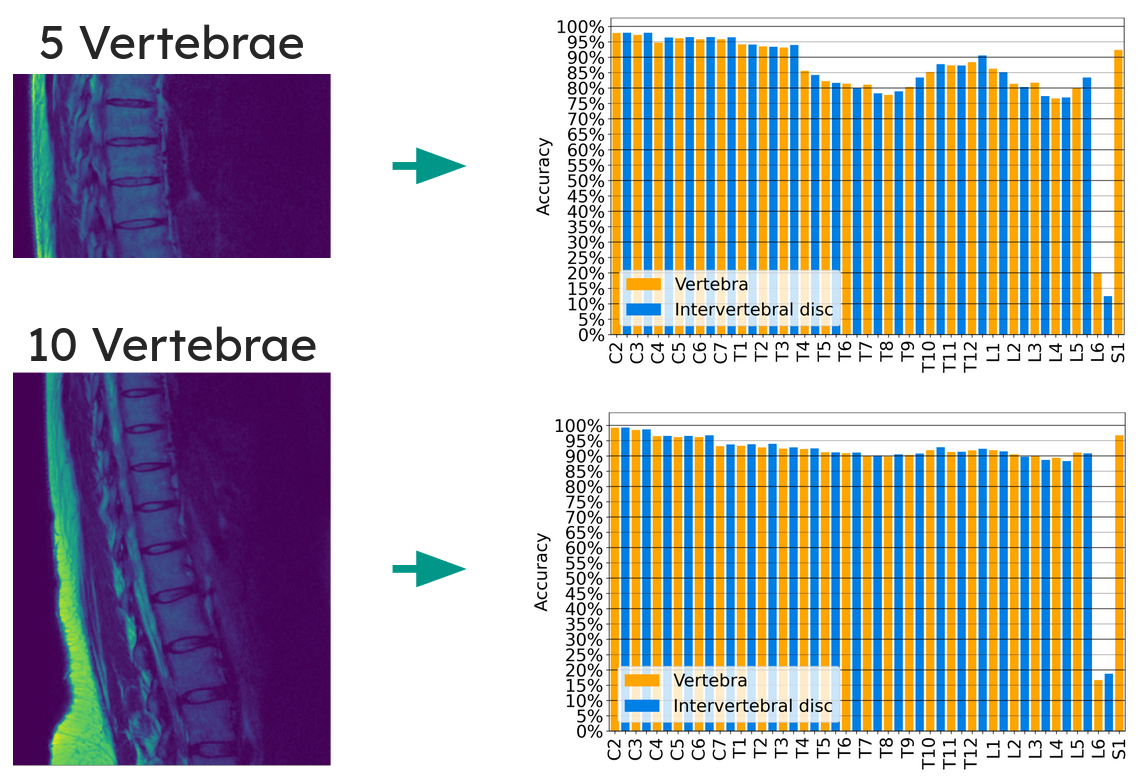
Per-vertebra accuracy for the two most challenging FOV sizes (5 and 10 visible vertebrae). Accuracy is fairly consistent across most vertebrae, with dips around the mid-thoracic region where vertebrae lack distinguishing features. C2 and S1 are easiest to identify due to their unique shapes. L6 and L6-S1 have notably poor accuracy because lumbarization only occurs in ~6.6% of the population, so the model rarely sees these classes during training.
Without the data generation pipeline (training only on the 162 manually annotated scans), accuracy drops by about 5 percentage points across all FOV sizes, confirming that the automatically generated training data from 10,833 scans is critical.
The pipeline also detected lumbarization in 6.6% and sacralization in 3.6% of the 10,833 patients, roughly consistent with literature (typically ~4.7% each).
Citation
If you use this work, please cite the master's thesis:
Brutenis Gliwa, "Anatomical Labeling of the Spine in Small Field-of-View MRI Scans",
Master's Thesis, University of Rostock, 2023.